Despite their size, tiny molecules store all of the information required to generate and maintain every bit of life on earth. All of the data needed to make a human lives in each of our cells. What does that tell you?
DNA is a pretty efficient data storage solution.
I covered a little bit of the potential for massive breakthroughs in data storage via DNA with the Shareable Science post on ‘Hachimoji’, an expanded genetic language. I also discussed it more in-depth in a Biotech 201 talk that you can watch here. So what’s new?
Microsoft has moved us one step closer toward using DNA data storage on a massive scale. Let’s look at how that happened.
Getting a computer to write its own DNA
One of the big barriers for successfully using DNA as an information storage solution is the time it takes to produce the coded DNA. Trust me, with plenty of experience at the Institute, I can assure you it’s complicated work! Many steps of the process have to be carried out by highly-trained scientists in a lab.
 If we want to store meaningful amounts of data in DNA, either we need a LOT more scientists (not saying I’m opposed…) or we need to find a way to automate more of the storage process.
If we want to store meaningful amounts of data in DNA, either we need a LOT more scientists (not saying I’m opposed…) or we need to find a way to automate more of the storage process.
That realm is where we find the most recent breakthrough.
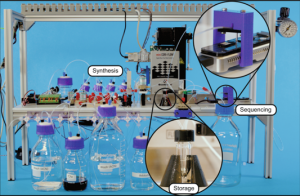
Microsoft engineers recently published a paper in Nature demonstrating an end-to-end automation of DNA storage. They created a system that used a small sequencer and several glass bottles to synthesize the DNA. In fact, the system coded, wrote and read a DNA sequence successfully.
All told, the whole system fit on a bench-top table and cost less than $10,000. In terms of genetic sequencing equipment, that’s a steal! Still, the system has some serious limitations that we’ll get to a little later.
The potential of DNA storage
Before we reel all the enthusiasm back in, let’s just talk about the incredible potential of DNA data storage for a minute.
If you followed the links in the opening paragraph, you’ve already seen some of the awesome power of a broad DNA data solution. But for those of you looking for the highpoints, take this from the Hachimoji Shareable Science post:
Humanity generated more data between 2015 and 2017 than in all of human history before it. DNA could help address that problem.
In 2017, scientists developed an approach that could store 215 petabytes (215 million gigabytes, if that gives you some perspective) in a single gram of DNA. That’s about the weight of a paperclip. Using that system, we could theoretically store all of humanity’s data — all of it, as in all of human history — in a container the size and weight of a pair of pickup trucks.
That’s a huge impact, but it only works if the process can be successfully automated. Still, it’s an exciting glimpse into the future, and it gives a sense of why we’re following this branch of technology through every step of its development.
The abundant limitations of current technology
Ok, ok, here’s where we’ve got to throw the wet blanket. Your next computer probably isn’t going be storing your video game saves or word processing documents on a strand of DNA.
 After all, the machine in the Microsoft paper? It coded, stored and read… one word. The test system wrote ‘Hello’. It took 21 hours.
After all, the machine in the Microsoft paper? It coded, stored and read… one word. The test system wrote ‘Hello’. It took 21 hours.
Ta da?
It’s exciting to imagine storing a warehouse’s worth of data—as Microsoft says—into a unit roughly the size of a set of Yahtzee dice. If that’s going to happen though, then the writing process would need to go literally millions of times faster.
The future of data storage
When you look at the massive drop in sequencing costs from the first genome to the genomes we sequence every day at the Institute, it’s obvious the technology is only getting better and more efficient.
One major hurdle for data storage remains automation, but it very much seems like we’re closer than we’ve ever been to understanding what an end-to-end automation process might look like.
Plenty of hurdles remain—speed, efficiency, accuracy, cost—but the potential and progress demonstrate that we’ve set off on an incredible course.
To schedule a media interview with Dr. Neil Lamb or to invite him to speak at an event or conference, please contact Margetta Thomas by email at mthomas@hudsonalpha.org or by phone: Office (256) 327-0425 | Cell (256) 937-8210
Get the Latest Sharable Science Delivered Straight to Your Inbox!
[gravityform id=19 title=false description=false ajax=true][wprpw_display_layout id=8]