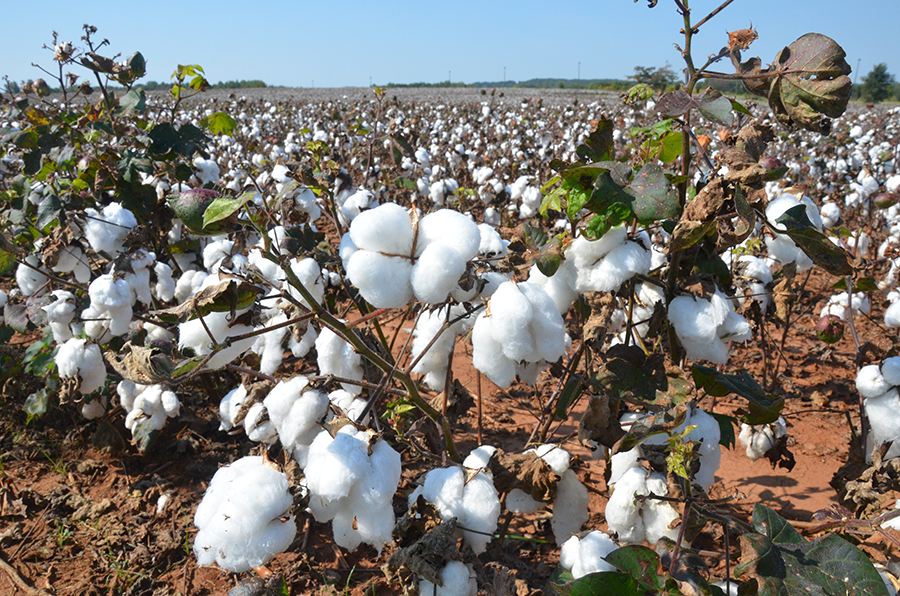
Faculty Investigator and Genomic Services Lab Director Shawn Levy, PhD, was a presenter at the 10X Genomics Advances in Genome Biology and Technology meeting in Orlando, Florida, where he discussed HudsonAlpha’s use of the Chromium system. Levy said that HudsonAlpha has been running the GemCode platform since around June and has received libraries generated by 10X Genomics on the Chromium that it has subsequently sequenced on Illumina’s HiSeq and HiSeq X Ten.
On average, the Chromium libraries have generated between 860 million and 884 million reads per lane and the genomes were sequenced to a mean depth of 32x to 34x with an insert size of 350 base pairs. PCR duplicates were between 5 percent and 6 percent and 99 percent of the SNPs called were phased in haplotype blocks with N50s averaging between 7 Mb and 9 Mb. The longest haplotype block was 30 Mb.
Levy said that the linked reads have made it easier to resolve structural variants. For instance, in one of the first genomes it sequenced, there was a copy number change that “wasn’t completely obvious from just the HiSeq” whole-genome sequencing data, but readily detectable with the linked reads from 10X Genomics.
Read the full story on genomeweb.